Major Steps Involved In Mechanism Of Protein Synthesis : Transcription And Translation
ADVERTISEMENTS:
Major steps involved in mechanism of Protein Synthesis are 1. Transcription and 2. Translation!
Biosynthesis of protein is under direct control of DNA in most cases or else under the control of genetic RNA where DNA is absent.
ADVERTISEMENTS:
Information for structure of a polypeptide is stored in a polynucleotide chain. In 1958 Crick proposed that the information present in DNA is transferred to RNA and then from RNA it is transferred to protein , and that this information does not flow in the reverse direction, that is, from protein to RNA to DNA.
DNA molecules provide the information for their own replication. This relationship between DNA, RNA and protein molecules is known as central dogma. Temin reported that retroviruses operate a central dogma reverse or feminism inside host cells.
Genomic RNA of these viruses first synthesizes DNA through reverse transcription this process is catalyzed by the enzyme reverse transcriptase, DNA then transfers information to messenger RNA which takes part in translation of the coded information to form polypeptide.
What Is Mrna Splicing
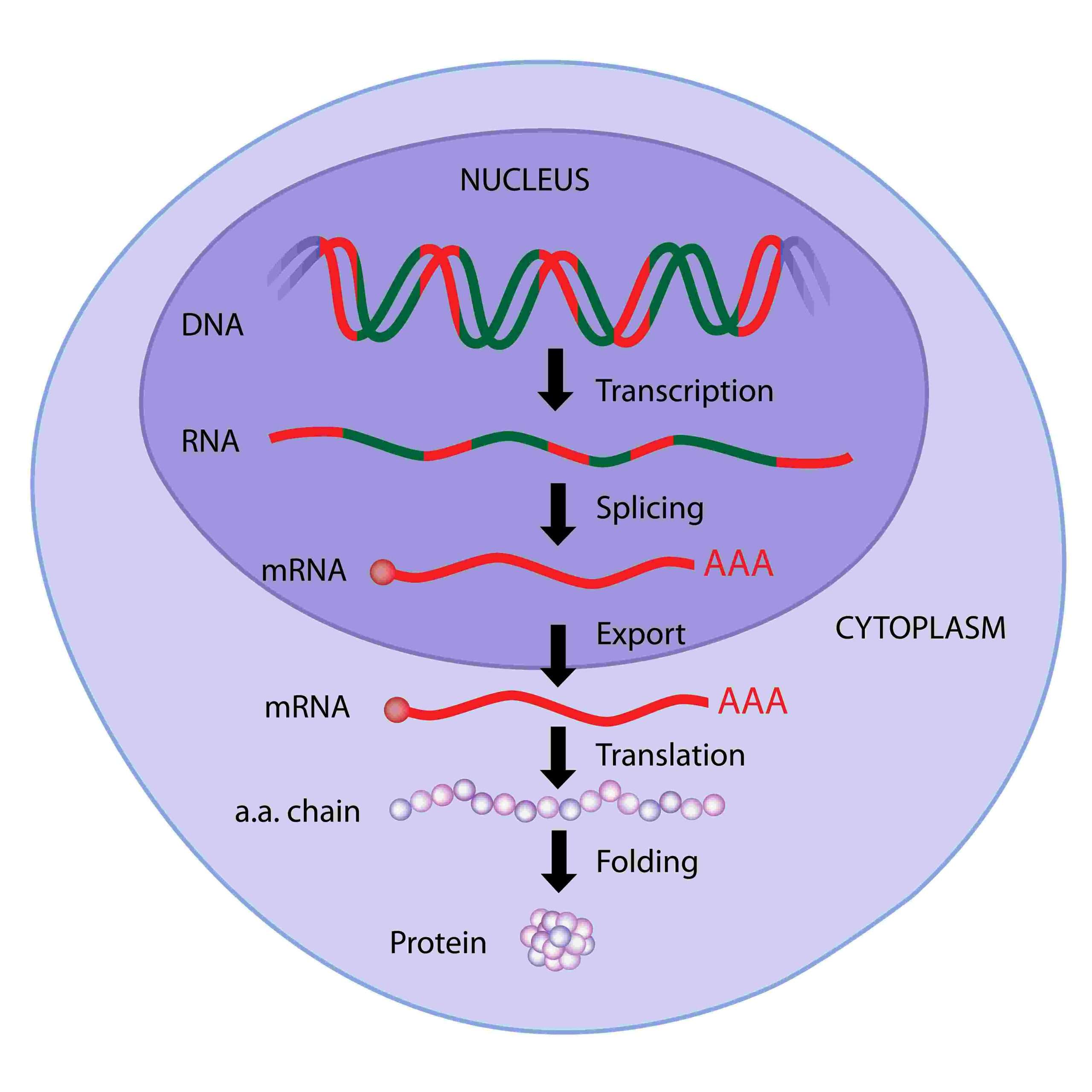
Eukaryotic cells contain introns and exons. But we only need the exons, as these are the coding regions. mRNA splicing describes the process of removing introns, so we have an mRNA strand containing just exons. Specialized enzymes called spliceosomes catalyse this process.
Fig. 2 – mRNA splicing
Once splicing is complete, the mRNA can diffuse out from the nuclear pore and towards the ribosome for translation.
Addition Of Complex Molecules
Post-translational modifications can incorporate more complex, large molecules into the folded protein structure. One common example of this is glycosylation, the addition of a polysaccharide molecule, which is widely considered to be most common post-translational modification.
In glycosylation, a polysaccharide molecule is covalently added to the target protein by glycosyltransferases enzymes and modified by glycosidases in the endoplasmic reticulum and Golgi apparatus. Glycosylation can have a critical role in determining the final, folded 3D structure of the target protein. In some cases glycosylation is necessary for correct folding. N-linked glycosylation promotes protein folding by increasing solubility and mediates the protein binding to protein chaperones. Chaperones are proteins responsible for folding and maintaining the structure of other proteins.
There are broadly two types of glycosylation, N-linked glycosylation and O-linked glycosylation. N-linked glycosylation starts in the endoplasmic reticulum with the addition of a precursor glycan. The precursor glycan is modified in the Golgi apparatus to produce complex glycan bound covalently to the nitrogen in an asparagine amino acid. In contrast, O-linked glycosylation is the sequential covalent addition of individual sugars onto the oxygen in the amino acids serine and threonine within the mature protein structure.
Don’t Miss: Garden Of Life Raw Organic Protein
Early Elongation Pauses Affect Protein Expression
Fig. 6: The identity of the first 5 amino acids impacts protein synthesis in a well-defined in vitro translation system.
a Thin-layer chromatography analysis of in vitro peptide synthesis using S35-labeled methionine . Sequences and GFP scores of tetra-peptides, penta-peptides, and hexa-peptides representing starts of wild type eGFP and two high expressing clones MVKYH and MVYKH are indicated. Protein synthesis was initiated from pelleted initiation complexes at time 0 and resolved over time . Points at 1, 5, 10, 20, 30, 45, 60, 90, 120, 180, 240, 300s are shown. Migration of tetra-peptide, penta-peptide, and hexa-peptide is indicated. Arrows indicate final hexa-peptide products of the reaction. b Analyses of accumulation of the tetra-peptide and hexa-peptide for MVSKGK, MVKYHK, and MVYKHK peptides. Amounts of radioactivity for tetra-peptide and hexa-peptide for the last three time points were normalized to total radioactivity and plotted in relation to MVSK or MVSKGK peptide amounts. Error bars represent standard deviation.
Also Check: Protein Shake Without Working Out
Ubiquitous Influence Of Codons 35 On Protein Synthesis
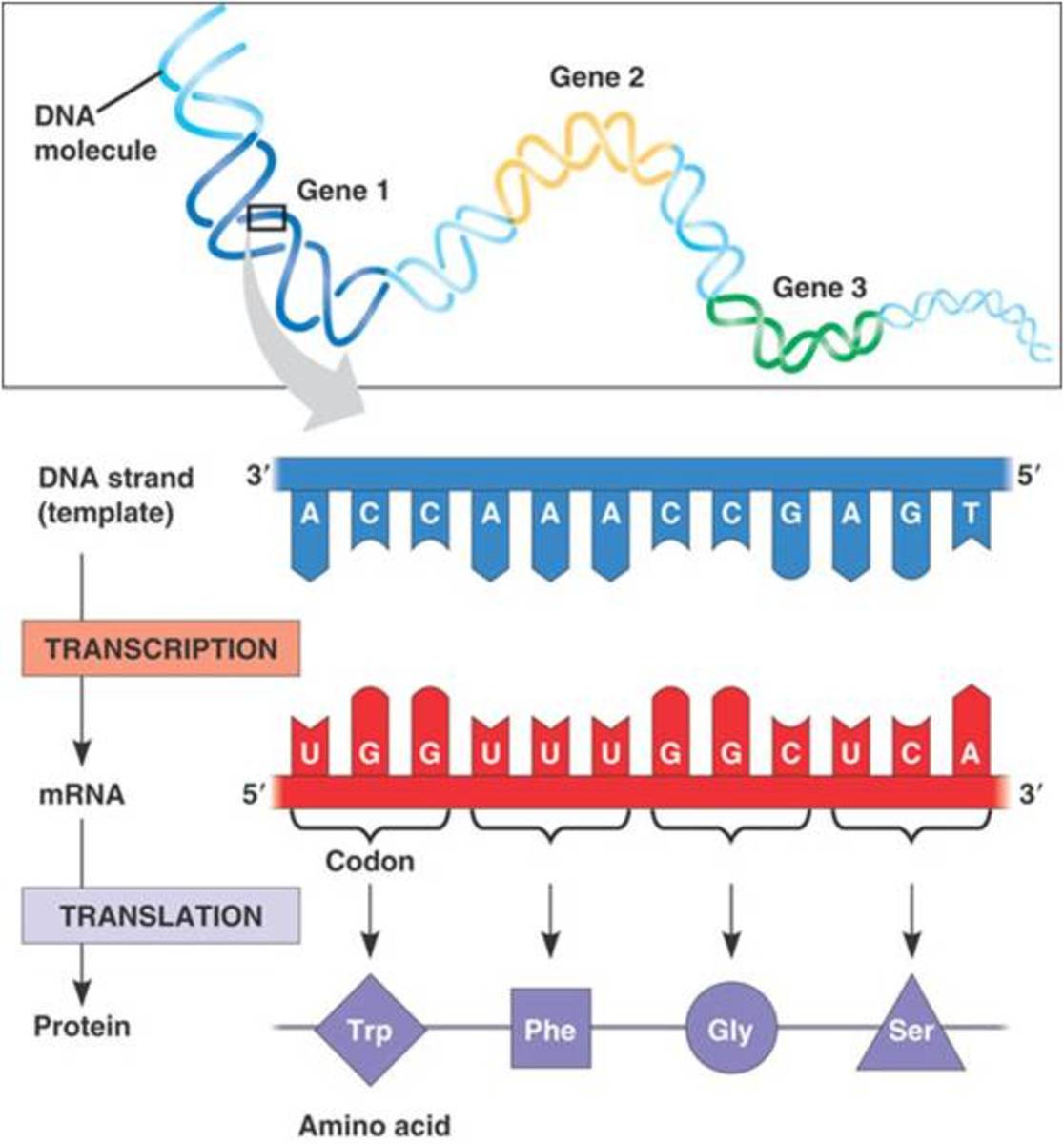
Fig. 5: The position and context of the motifs around the initiation codon is critical for protein-synthesis yield.
To further examine the effects of codons 35 on protein expression and impact of the 5 non-coding sequences we turned towards vectors that express polycistronic mRNAs. The majority of the E. coli genes are expressed from polycistronic operons where expression of each individual transcript is dependent on ORF-centric structures. We tested expression of the same eGFP variants as in previous experiments, however, now cloned as the second ORF in polycistronic operon with mCherry into two different vectors . In addition, the two vectors had different promoter, intergenic and ribosome binding sequences close to the translation start site. Expression of mCherry reporter was uniform for all constructs regardless of the vector, while eGFP variants were expressed at different levels that correlated with the GFP score determined from our library . Noticeably in vivo expression of eGFP variants from both polycistronic vectors, as well as in vitro expression from T7 polymerase transcribed polycistronic mRNAs, recapitulated data using monocistronic reporter . As such, previously observed differences between eGFP variants were maintained regardless of the polycistronic arrangement, differences in promoter, intergenic sequences or RBS.
Recommended Reading: Best Protein For Hair Growth
What Four Organelles Are Involved In Protein Synthesis
Cell organelles that participate in protein synthesis are golgi bodies, ribosomes and endoplasmic reticulum.
Which organelle is important for protein synthesis?
RibosomesRibosomes on the outer surface of the endoplasmic reticulum play an important role in protein synthesis within cells.
What organelle is responsible for protein building?
ribosomeA ribosome is an organelle that serves as the site of protein synthesis. It is composed of two ribosomal RNA subunits that wrap around mRNA to start the process of translation, followed by protein synthesis.
What is the list of organelles that take part in protein synthesis quizlet?
What is the list of organelles that take part in protein synthesis? Nucleus has instructions for making proteins Nucleolus makes ribosomes Ribosomes make proteins ER transports proteins within cell Golgi packages proteins which can then be exported through the cell membrane.
Translocation Of The Protein Molecule:
Two types of polyribosome shave been discovered that are involved in this process:
- Free polyribosomes
- Membrane-bound polyribosomes.
Upon termination of protein synthesis in the free ribosome, the prepared ribosome releases the protein into the cytoplasm. Special types of processes are used to transport some of these specialised proteins to the mitochondria and nucleus.
In membrane-bound polyribosomes, on the other hand, a polypeptide chain that develops on mRNA is introduced into the ER membranes lumen. Some of the proteins also compose parts of the membrane structure.
Even yet, only a few proteins are released into the lumen and integrated into Golgi body vesicles. They can also change the protein through glycosylation, which is the addition of sugar residues. As a result, the vesicles shape a bond with the plasma
membrane and the proteins are sooner or later released.
Don’t Miss: High Protein Low Calorie Meals
How Are The Vesicles Involved In Protein Synthesis
The vesicles from the Golgi merge with the cell membrane, and the proteins within them can either become integrated into the membrane or be secreted. The first step in protein production is the transcription of DNA to mRNA in the nucleus. The next organelles involved are ribosomes.
What are the steps in the protein synthesis process?
Protein synthesis is the process in which cells make proteins. It occurs in two stages: transcription and translation. Transcription is the transfer of genetic instructions in DNA to mRNA in the nucleus. It includes the steps of initiation, elongation, and termination.
https://www.youtube.com/watch?v=26y1PCkWiIc
Protein Synthesis Steps In Brief
The process of protein synthesis translates the codons of the messenger RNA into the 20-symbol code of amino acids that build the polypeptide chain of the proteins. The process of mRNA translation begins from its 5-end towards its 3-end as the polypeptide chain is synthesized from its amino-terminal to its carboxyl-terminal . There are almost no significant differences in the protein synthesis steps in prokaryotes and eukaryotes, however there is one major distinction between the structure of the mRNAs prokaryotes often have several coding regions , while the eukaryotic mRNA has only one coding region .
The main protein synthesis steps are:
In most of the aspects, the process in eukaryotes follow the same simple protein synthesis steps as in prokaryotes. However there are specific differences that could be outlined. For example, one important difference is that in prokaryotic cells the process of translation starts before transcription is completed. This coupling is defined because prokaryotes have no nuclear membrane and thus there is no physical separation of the two processes.
Don’t Miss: Before And After 7 Day Protein Shake Diet Results
Regulating Eif4eeif4g Interactions By 4e
In higher eukaryotes, 4E-BPs play a prominent role in controlling eIF4E function and cellular translation . The 4E-BPs are a diverse set of proteins that share a common, albeit rather degenerate, motif YxxxxL that enables them to compete with eIF4G for binding to the surface of the cap-binding protein eIF4E. The 4E-BPs regulate a variety of cellular and developmental processes in higher eukaryotes . Yeast has two characterized 4E-BPs that contain the consensus eIF4E-binding motif, Caf20 and Eap1, which are 18 and 70 kDa, respectively . Caf20 and Eap1 share no sequence similarity outside the eIF4E-binding motif, mutation of which abrogates eIF4E interactions . Because mutations in eIF4E have differential impacts on the binding of eIF4G and Caf20, the eIF4E-interaction interfaces with eIF4G and Caf20 are likely overlapping, but distinct . This potentially enables Caf20 to displace eIF4G from eIF4E. Recent structural studies show higher eukaryote 4E-BPs share similar eIF4E-binding properties . These data suggest the yeast proteins are parallels of the higher eukaryote 4E-BPs.
Differences Between Eukaryotic And Prokaryotic Protein Synthesis
The overall scheme of protein synthesis is similar in all living cells. However, there are significant differences between bacteria and eukaryotes. These are summarized in Table 13.04 and discussed in the following sections. Note that eukaryotic cells contain mitochondria and chloroplasts, which have their own DNA and their own ribosomes. The ribosomes of these organelles operate similarly to those of bacteria and will be considered separately below. In eukaryotic protein synthesis, it is usually the cytoplasmic ribosomes that translate nuclear genes. Several aspects of eukaryotic protein synthesis are more complex. The ribosomes of eukaryotic cells are larger and contain more rRNA and protein molecules than those of prokaryotes. In addition, eukaryotes have more initiation factors and a more complex initiation procedure.
Table 13.04. Comparison of Protein Synthesis
Eukaryotic ribosomes are larger and more complex than those of prokaryotes.
A few aspects of protein synthesis are actually less complex in eukaryotes. In prokaryotes, mRNA is polycistronic and may carry several genes that are translated to give several proteins. In eukaryotes, each mRNA is monocistronic and carries only a single gene, which is translated into a single protein. In prokaryotes, the genome and the ribosomes are both in the cytoplasm, whereas in eukaryotes the genome is in the nucleus. Consequently, coupled transcription and translation is not possible for eukaryotes .
Internal Ribosome Entry Sites
Also Check: Which Protein Powder Is Best For Muscle Gain
Translation: Making Protein Synthesis Possible
- B.A., Biology, Emory University
- A.S., Nursing, Chattahoochee Technical College
Protein synthesis is accomplished through a process called translation. After DNA is transcribed into a messenger RNA molecule during transcription, the mRNA must be translated to produce a protein. In translation, mRNA along with transfer RNA and ribosomes work together to produce proteins.
Protein Synthesis And Genetic Code
A genetic code or codon is a trinucleotide sequence that identifies a particular amino acid. A codon is a group of three nucleotides so it is called a trinucleotide sequence. For example, a sequence GCC codes for the particular amino acid alanine.
Similarly, 64 codons are present that encode different amino acids. These codons are present in the mRNA template that adjuncts to anticodons present in the tRNA molecule. These anticodons are also trinucleotide sequences.
mRNA, tRNA, and rRNA
There are three types of RNA involved in the process of protein synthesis. The first type is mRNA which carries codons that are converted into an amino acid chain. These mRNA molecules are produced by a DNA template in the nucleus of the cell.
However, in prokaryotes, it synthesizes in the cytoplasm. The mRNA is consists of a 5 cap, 5 UTR region, coding region, 3 UTR region, and a poly chain. The coding region of mRNA is responsible for gene expression which has a start codon at 5 end and a stop codon at 3 end.
Another type of RNA is the tRNA molecule, also called transfer RNA. The transfer of particular amino acid to the ribosome is completed by a tRNA molecule. It has a cloverleaf resembling structure that has two major sites, named as- anticodon arm and accepter stem.
The anticodons are present in the anticodon arm and the accepter stem specifies the particular amino acid that needs to be attached. Now the ribosome translates the mRNA template into a polypeptide chain.
Initiation
Read Also: High Protein Diet Constipation Relief
Formation Of Covalent Bonds
Many proteins produced within the cell are secreted outside the cell to function as extracellular proteins. Extracellular proteins are exposed to a wide variety of conditions. In order to stabilize the 3D protein structure, covalent bonds are formed either within the protein or between the different polypeptide chains in the quaternary structure. The most prevalent type is a disulfide bond . A disulfide bond is formed between two cysteine amino acids using their side chain chemical groups containing a Sulphur atom, these chemical groups are known as thiol functional groups. Disulfide bonds act to stabilize the pre-existing structure of the protein. Disulfide bonds are formed in an oxidation reaction between two thiol groups and therefore, need an oxidizing environment to react. As a result, disulfide bonds are typically formed in the oxidizing environment of the endoplasmic reticulum catalyzed by enzymes called protein disulfide isomerases. Disulfide bonds are rarely formed in the cytoplasm as it is a reducing environment.
Study Reveals Key Step In Protein Synthesis
Protein-building ribosome caught in the act, showing a crucial mechanism in the translation of genetic code into protein structures
This image of the molecules involved in translation of genes into proteins shows the ribosome in transparent rendering. In the foreground are the messenger RNA , the elongation factor EF-G , and the four sequential positions of transfer RNA as it moves from right to left during translocation . The tRNA positions are like four frames of a molecular movie describing its movement through the ribosome during protein synthesis.
Scientists at the University of California, Santa Cruz, have trapped the ribosome, a protein-building molecular machine essential to all life, in a key transitional state that has long eluded researchers. Now, for the first time, scientists can see how the ribosome performs the precise mechanical movements needed to translate genetic code into proteins without making mistakes.
This is something that the whole field has been pursuing for the past decade, said Harry Noller, Sinsheimer Professor of Molecular Biology at UC Santa Cruz. Weve trapped the ribosome in the middle of its movement during translocation, which is the most interesting, profound, and complex thing the ribosome does.
Noller has spent decades working to understand how the ribosome works. Being able to see how it moves, he said, is an exciting moment.
Read Also: Think High Protein Bars Review
Enzymes Involved In Transcription
DNA helicase is the enzyme responsible for the early step of unwinding and unzipping. This enzyme catalyses the breaking of the hydrogen bonds found between complementary base pairs and allows the template strand to be exposed for the next enzyme, RNA polymerase.
RNA polymerase travels along the strand and catalyses the formation of phosphodiester bonds between adjacent RNAnucleotides. Adenine pairs with uracil, while cytosine pairs with guanine.
Remember: in RNA, adenine pairs with uracil. In DNA, adenine pairs with thymine.
Site Of Protein Synthesis
The site of protein synthesis is twofold. Transcription occurs within the cell nucleus where DNA is located. Once the mRNA copy of a small section of DNA has been made it travels through the nuclear pores and into the cell cytoplasm. In the cytoplasm, the strand of mRNA will move towards a free ribosome or one attached to the rough endoplasmic reticulum. Then the next step of protein synthesis translation can begin.
You May Like: Best Plant Base Protein Powder
Activation Of Amino Acids:
Thereaction is brought about when amino acids come to interact with ATP molecules catalyzed by aminoacyl RNA synthetase. The aminoacyl AMP enzyme complex is generated as a result of the reaction between amino acid and adenosine triphosphate , which is mediated by the aforementioned enzyme. The complex is as follows:
AA + ATP Enzyme -AA AMP enzyme complex + PP
Image showing DNA transcription to mRNA
Its worth noting that different amino acids require different aminoacyl RNA synthetases.
Initiation Of A Polypeptide Chain:
The ribosome accepts charged tRNA. In all organisms, protein synthesis occurs in the ribosome that is normally attached to the Golgi bodies in the cytoplasm. The SOS subunit of the 70S type ribosome interacts with the mRNA. Ribosomes are small complex molecules, responsible for protein synthesis and made up of 2 components- rRNA and proteins. Ribosomes also catalyze the creation of peptide bonds . Ribosomes are classified into two types: large and tiny.
Scientists represent each amino acid by three nucleic acid sequences known as codons. Based on the arrangement of the nitrogenous bases, this information is present in the mRNA. The amino acid methionine is transcribed as an initiating codon by the codon AUG but rarely by GUG , which is always responsible for starting polypeptide chains in prokaryotes. In prokaryotes, the formation of the starting amino acid methionine is a must.
Illustration showing translation process with the cycle of tRNA codon-anti-codon pairing and amino acid incorporation into the growing polypeptide chain by the ribosome
Ribosomes have two binding sites for amino-acyl-tRNA.
Don’t Miss: What Is An Incomplete Protein