Study Reveals Key Step In Protein Synthesis
Protein-building ribosome caught in the act, showing a crucial mechanism in the translation of genetic code into protein structures
This image of the molecules involved in translation of genes into proteins shows the ribosome in transparent rendering. In the foreground are the messenger RNA , the elongation factor EF-G , and the four sequential positions of transfer RNA as it moves from right to left during translocation . The tRNA positions are like four frames of a molecular movie describing its movement through the ribosome during protein synthesis.
Scientists at the University of California, Santa Cruz, have trapped the ribosome, a protein-building molecular machine essential to all life, in a key transitional state that has long eluded researchers. Now, for the first time, scientists can see how the ribosome performs the precise mechanical movements needed to translate genetic code into proteins without making mistakes.
“This is something that the whole field has been pursuing for the past decade,” said Harry Noller, Sinsheimer Professor of Molecular Biology at UC Santa Cruz. “We’ve trapped the ribosome in the middle of its movement during translocation, which is the most interesting, profound, and complex thing the ribosome does.”
Noller has spent decades working to understand how the ribosome works. Being able to see how it moves, he said, is an exciting moment.
S In Protein Synthesis
| BIOCHEMISTRY 3 |
|
5 steps in protein synthesis |
|
|
Addition Of Chemical Groups
Following translation, small chemical groups can be added onto amino acids within the mature protein structure. Examples of processes which add chemical groups to the target protein include methylation, acetylation and phosphorylation.
Methylation is the reversible addition of a methyl group onto an amino acid catalyzed by methyltransferase enzymes. Methylation occurs on at least 9 of the 20 common amino acids, however, it mainly occurs on the amino acids lysine and arginine. One example of a protein which is commonly methylated is a histone. Histones are proteins found in the nucleus of the cell. DNA is tightly wrapped round histones and held in place by other proteins and interactions between negative charges in the DNA and positive charges on the histone. A highly specific pattern of amino acid methylation on the histone proteins is used to determine which regions of DNA are tightly wound and unable to be transcribed and which regions are loosely wound and able to be transcribed.
Don’t Miss: What Does Protein Do In Your Body
Protein Synthesis Video Animation
Read Also:
Antimicrobial agents are used as protein synthesis inhibitors which include:
Process Of Protein Synthesis In Eukaryotes:
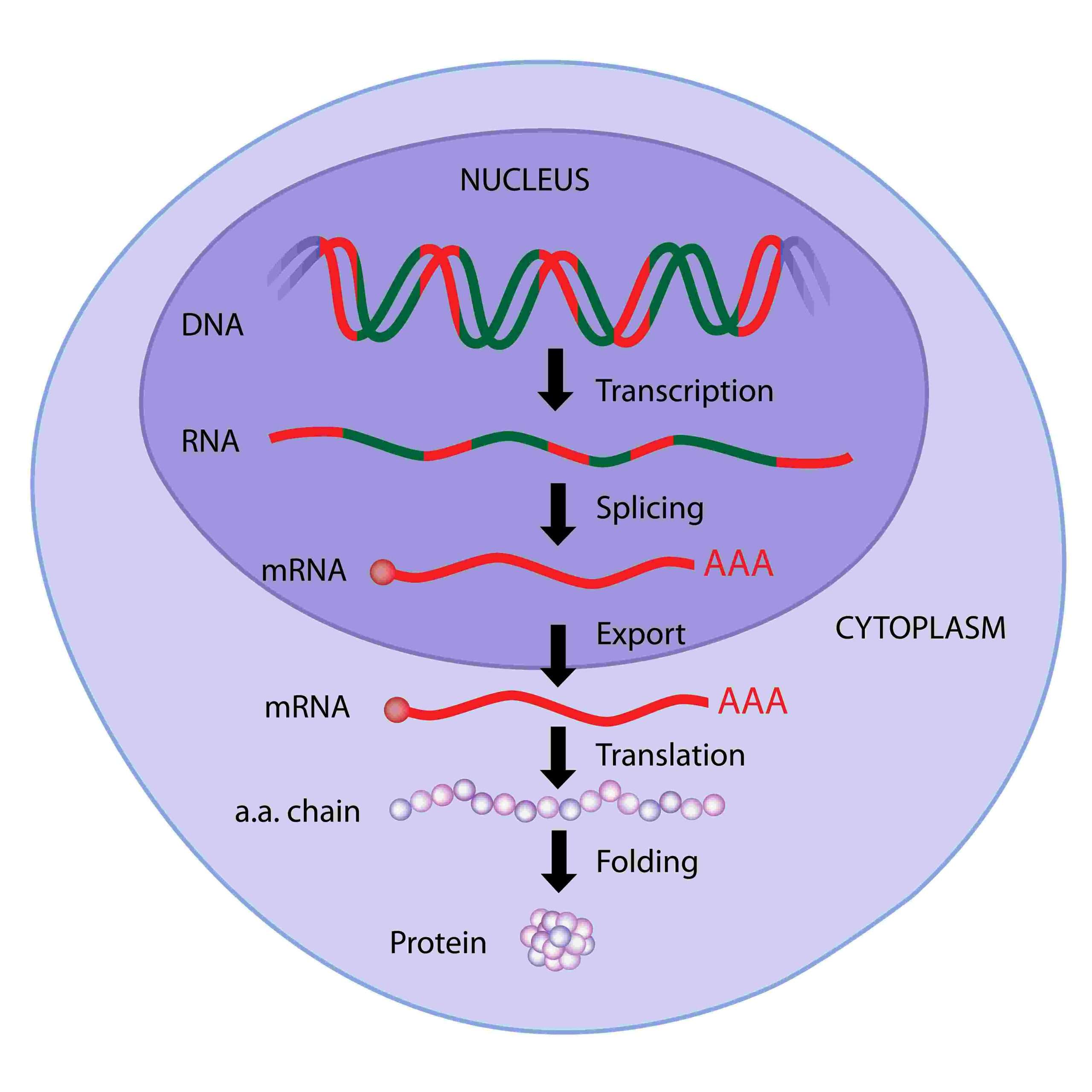
Protein synthesis in eukaryotes is a bit more complicated due to the presence of introns in the RNA that is transcribed. Hence the introns must be removed by the transcriptional process of splicing to ligate only the exons and produce mRNA which can be translated into protein.
Biochemistry of RNA splicing Image: Wikipedia
Post-transcriptional processing consists of a total of 4 steps in eukaryotic hnRNA:
Read Also: Sascha Fitness Hydrolyzed Whey Protein Isolate
Show The People Behind The Science
Storytelling is the most helpful method to grab students attention. Therefore, I spent some time digging a little history of the process to help students relate with the discoverer.
It wont be fair to crown the discovery of protein synthesis to a particular scientist. Zamecnik, a nester of protein synthesis, stated that an impressive list of pioneers blazed the trail to the present scene, making it an eminently multidisciplinary endeavor.
This scientific endeavor dates back to the beginning of the 20th century when Franz Hofmeister and Emil Fischer recognized the peptide bond structure of proteins. Fritz Lipmann enlightened us about the presence of a high-energy phosphate intermediate, while Max Bergmann reported the specificity of proteolytic enzymes. Torbjörn Caspersson and Jean Brachet unraveled the role of RNA in protein synthesis, followed by the discovery of Frederick Sanger, who discovered the first primary protein structure. Sanger also showed the amino acid specification to produce insulin. All this work led up to the efforts of George Palade, who presented visual evidence of protein synthesis with particulate structures in the cytoplasm acting as the cellular sites.
Early Elongation Pauses Affect Protein Expression
Fig. 6: The identity of the first 5 amino acids impacts protein synthesis in a well-defined in vitro translation system.
a Thin-layer chromatography analysis of in vitro peptide synthesis using S35-labeled methionine . Sequences and GFP scores of tetra-peptides, penta-peptides, and hexa-peptides representing starts of wild type eGFP and two high expressing clones MVKYH and MVYKH are indicated. Protein synthesis was initiated from pelleted initiation complexes at time 0 and resolved over time . Points at 1, 5, 10, 20, 30, 45, 60, 90, 120, 180, 240, 300s are shown. Migration of tetra-peptide, penta-peptide, and hexa-peptide is indicated. Arrows indicate final hexa-peptide products of the reaction. b Analyses of accumulation of the tetra-peptide and hexa-peptide for MVSKGK, MVKYHK, and MVYKHK peptides. Amounts of radioactivity for tetra-peptide and hexa-peptide for the last three time points were normalized to total radioactivity and plotted in relation to MVSK or MVSKGK peptide amounts. Error bars represent standard deviation.
Also Check: Protein Shake Without Working Out
The Mechanism Of Protein Synthesis
As with mRNA synthesis, protein synthesis can be divided into three phases: initiation, elongation, and termination. The process of translation is similar in prokaryotes and eukaryotes. Here well explore how translation occurs in E. coli, a representative prokaryote, and specify any differences between prokaryotic and eukaryotic translation.
Protein Synthesis Enzymes And Functions
- Peptidyl transferase is the main enzyme used in Translation. It is found in the ribosomes with an enzymatic activity that catalyzes the formation of a covalent peptide bond between the adjacent amino acids.
- The enzymes activity is to form peptide bonds between adjacent amino acids using tRNAs during translation.
- The enzymes activity uses two substrates of which one has the growing peptide chain and the other bears the amino acid that is added to the chain.
- It is located in the large subunit of the ribosomes and therefore, the primary function of peptidyl transferase is to catalyze the addition of amino acid residues allowing the polypeptide chain to grow.
- The peptidyl transferase enzyme is entirely made up of RNA and its mechanism is mediated by ribosomal RNA , which is a ribozyme, made up of ribonucleotides.
- In prokaryotes, the 23S subunit contains the peptidyl transferase between the A-site and the O-site of tRNA while in eukaryotes, it is found in the 28S subunit.
You May Like: Levels Of Structure In Proteins
Macrolides Chloramphenicol And Oxazolidinones
Protein synthesis, that is, the process of mRNA translation, consists of three steps . These steps involve ribosomes and several other cytoplasmic factors, which play important roles during this process. The ribosome is a complex organelle and is composed of two subunits, the 50 S and 30 S units. Together, they organize the formation of a complex between f-Met-charged-aminoacyl-tRNA and mRNA transcript. This process involves several other initiation factors. Aminoglycosides and tetracycline are one of the most diverse classes of antibiotics , which target this machinery and inhibit protein synthesis in bacteria. Based on the target they choose for inhibitioneither the 50 S inhibitors and the 30 S inhibitorsthey are subdivided into two classes of these antibiotics .
50 S ribosome inhibitors include the macrolide , lincosamide , streptogramin , amphenicol , and oxazolidinone classes of antibiotics. Generally, 50 S ribosome inhibitors bind with it and block either protein translation initiation or peptidyl-tRNAs translocation. This is caused by the inhibition of the peptidyltransferase reaction which is responsible for the elongation of emerging peptide chains .
Alexei A. Sharov, in, 2018
Initiation Of A Polypeptide Chain:
The ribosome accepts charged tRNA. In all organisms, protein synthesis occurs in the ribosome that is normally attached to the Golgi bodies in the cytoplasm. The SOS subunit of the 70S type ribosome interacts with the mRNA. Ribosomes are small complex molecules, responsible for protein synthesis and made up of 2 components- rRNA and proteins. Ribosomes also catalyze the creation of peptide bonds . Ribosomes are classified into two types: large and tiny.
Scientists represent each amino acid by three nucleic acid sequences known as codons. Based on the arrangement of the nitrogenous bases, this information is present in the mRNA. The amino acid methionine is transcribed as an initiating codon by the codon AUG but rarely by GUG , which is always responsible for starting polypeptide chains in prokaryotes. In prokaryotes, the formation of the starting amino acid methionine is a must.
Illustration showing translation process with the cycle of tRNA codon-anti-codon pairing and amino acid incorporation into the growing polypeptide chain by the ribosome
Ribosomes have two binding sites for amino-acyl-tRNA.
Read Also: High C Reactive Protein Means
Translation Process In Protein Synthesis
During the translation process, the small and large subunits of a ribosome close over a strand of mRNA, trapping it loosely inside. Ribosomes arrange the strand into codons or sets of three nitrogenous base letters. This is because the code for a single amino acid the most basic form of a protein is a three-letter nucleobase code.
As ribosomes recognize parts of code, we can say they understand it. The jumble of copied letters made during the transcription phase can be read and understood in the translation phase.
For example, GGU, GGC, GGA, and GGG code for the amino acid known as glycine. Most amino acids have multiple codes as this lowers the chance of mistakes if RNA polymerase accidently connects adenine instead of cytosine to GG, it doesnt matter. Both GGC and GGA code for the same amino acid. You can see a list of mRNA codons for the twenty non-essential amino acids here.
There is only one start codon code AUG. Three codons TAA, TAG, and TGA represent stop codons. Neither start nor stop codons match the code for an amino acid they are non-coding. The single start and three stop codons are clearly marked on this codon wheel.
As with the process of transcription, translation within the ribosome is also split into the three stages of initiation, elongation, and termination.
Activation Of Amino Acids:
Thereaction is brought about when amino acids come to interact with ATP molecules catalyzed by aminoacyl RNA synthetase. The aminoacyl AMP enzyme complex is generated as a result of the reaction between amino acid and adenosine triphosphate , which is mediated by the aforementioned enzyme. The complex is as follows:
AA + ATP Enzyme -AA AMP enzyme complex + PP
Image showing DNA transcription to mRNA
Its worth noting that different amino acids require different aminoacyl RNA synthetases.
You May Like: Should I Drink Protein Shakes Everyday
Iiihnovel Functions Of Aarss
In addition to their critical role in protein synthesis, it has become clear that AARSs are involved in several other cellular pathways. Some AARSs regulate their own transcription and translation, while others contribute to splicing activities in mitochondria. Nuclear aminoacylation of tRNAs by imported AARSs is thought to be a quality control mechanism to ensure that only mature, fully active tRNAs are released efficiently to the cytoplasm for protein synthesis. Programmed cell death also appears to have an AARS componenthuman tyrosyltRNA synthetase can be proteolytically cleaved into two polypeptides with distinct cytokine activities, despite the lack of such activity in the full-length TyrRS. It is likely that in time many more nontranslational functions of AARS will be identified.
Neha Upmanyu, Viveka Nand Malviya, in, 2020
Protein Synthesis And Genetic Code
A genetic code or codon is a trinucleotide sequence that identifies a particular amino acid. A codon is a group of three nucleotides so it is called a trinucleotide sequence. For example, a sequence GCC codes for the particular amino acid alanine.
Similarly, 64 codons are present that encode different amino acids. These codons are present in the mRNA template that adjuncts to anticodons present in the tRNA molecule. These anticodons are also trinucleotide sequences.
mRNA, tRNA, and rRNA
There are three types of RNA involved in the process of protein synthesis. The first type is mRNA which carries codons that are converted into an amino acid chain. These mRNA molecules are produced by a DNA template in the nucleus of the cell.
However, in prokaryotes, it synthesizes in the cytoplasm. The mRNA is consists of a 5 cap, 5 UTR region, coding region, 3 UTR region, and a poly chain. The coding region of mRNA is responsible for gene expression which has a start codon at 5 end and a stop codon at 3 end.
Another type of RNA is the tRNA molecule, also called transfer RNA. The transfer of particular amino acid to the ribosome is completed by a tRNA molecule. It has a cloverleaf resembling structure that has two major sites, named as- anticodon arm and accepter stem.
The anticodons are present in the anticodon arm and the accepter stem specifies the particular amino acid that needs to be attached. Now the ribosome translates the mRNA template into a polypeptide chain.
Initiation
Don’t Miss: Best Whey Protein For Beginners
Explain Which Rna Is Involved In Protein Synthesis
The process of translation takes place in the cytoplasm of the self Nucleus undergoes some modifications before it gets translated. Removal of unneeded sections and modification is done to protect Me ends °Mlle mRNA molecule and our.The RNA molecule is ready for translation protein synthesis. Another RNA involved is called transfer RNA is in the shape of a cloverleaf with 3 loops. It has an anticodon site that has a special section in the middle of the loop and it also contains an amino acid site at its one end. Anticodon plays an important role to recognize a specific messenger RNA called start codon.
Ubiquitous Influence Of Codons 35 On Protein Synthesis
Fig. 5: The position and context of the motifs around the initiation codon is critical for protein-synthesis yield.
To further examine the effects of codons 35 on protein expression and impact of the 5 non-coding sequences we turned towards vectors that express polycistronic mRNAs. The majority of the E. coli genes are expressed from polycistronic operons where expression of each individual transcript is dependent on ORF-centric structures. We tested expression of the same eGFP variants as in previous experiments, however, now cloned as the second ORF in polycistronic operon with mCherry into two different vectors . In addition, the two vectors had different promoter, intergenic and ribosome binding sequences close to the translation start site. Expression of mCherry reporter was uniform for all constructs regardless of the vector, while eGFP variants were expressed at different levels that correlated with the GFP score determined from our library . Noticeably in vivo expression of eGFP variants from both polycistronic vectors, as well as in vitro expression from T7 polymerase transcribed polycistronic mRNAs, recapitulated data using monocistronic reporter . As such, previously observed differences between eGFP variants were maintained regardless of the polycistronic arrangement, differences in promoter, intergenic sequences or RBS.
Read Also: Soy Protein Vs Pea Protein