S Involved In Protein Synthesis Mechanism
ADVERTISEMENTS:
The mechanism of protein synthesis has been studied thoroughly in E.coli. The process of protein synthesis on 70S ribosomes is described in detail below. The summary of the various steps in the mechanism of protein synthesis is shown in .
Step 1 Transcription:
One strand of DNA molecule functions as a template for the formation of mRNA. This mRNA contains the message for the protein to be synthesised. As soon as the mRNA is formed, it leaves the nucleus and reaches the cytoplasm where it attaches with the 30S subunit of ribosomes.
Step 2 Attachment of mRNA to 30S Subunit of Ribosomes:
ADVERTISEMENTS:
In prokaryotic cells, ribosomes are found in a dissociated and an inactive state. The mRNA binds to the 30S subunit. Specific sequences of 16 S rRNA of the small subunit of the ribosome binds to complementary sequence of mRNA. The first amino acid, which is usually N formyl methionine-tRNA attaches to the mRNA to form the initiation complex.
This process is aided by GTP and three protein factors . The ribosome is uniquely designed to bring mRNA and tRNA together. The mRNA is threaded through the small subunit tunnel, while the large and subunit fit together to form tRNA binding pockets. After the formation of the initiation complex, the 50S subunit joins with the smaller subunit to form the 70S ribosome.
Step 3 Transfer of Amino Acids to the Site of Protein Synthesis:
Step 4 Initiation of Protein Synthesis:
Step 5 Elongation of Polypeptide Chain:
Many Proteins Are Controlled By Regulated Destruction
One function of intracellular proteolytic mechanisms is to recognize and eliminate misfolded or otherwise abnormal proteins, as just described. Yet another function of these proteolytic pathways is to confer short half-lives on specific normal proteins whose concentrations must change promptly with alterations in the state of a cell. Some of these short-lived proteins are degraded rapidly at all times, while many others are conditionally short-lived, that is, they are metabolically stable under some conditions, but become unstable upon a change in the cell’s state. For example, mitotic cyclins are long-lived throughout the cell cycle until their sudden degradation at the end of , as explained in Chapter 17.
How is such a regulated destruction of a controlled? A variety of mechanisms are known, as illustrated through specific examples later in this book. In one general class of mechanism , the activity of a is turned on either by E3 or by an allosteric transition in an E3 protein caused by its binding to a specific small or large . For example, the is a multisubunit ubiquitin ligase that is activated by a cell-cycle-timed addition at . The activated APC then causes the degradation of mitotic cyclins and several other regulators of the -anaphase transition .
The Organization Of Mrnas And The Initiation Of Translation
Although the mechanisms of protein synthesis in prokaryotic and are similar, there are also differences, particularly in the signals that determine the positions at which synthesis of a chain is initiated on an mRNA template . Translation does not simply begin at the 5´ end of the mRNA it starts at specific initiation sites. The 5´ terminal portions of both prokaryotic and eukaryotic mRNAs are therefore noncoding sequences, referred to as 5´untranslated regions. Eukaryotic mRNAs usually encode only a single polypeptide chain, but many prokaryotic mRNAs encode multiple polypeptides that are synthesized independently from distinct initiation sites. For example, the E. coli lac consists of three genes that are translated from the same mRNA . Messenger RNAs that encode multiple polypeptides are called , whereas mRNAs encode a single polypeptide chain. Finally, both prokaryotic and eukaryotic mRNAs end in noncoding 3´untranslated regions.
Prokaryotic and eukaryotic mRNAs. Both prokaryotic and eukaryotic mRNAs contain untranslated regions at their 5´ and 3´ ends. Eukaryotic mRNAs also contain 5´ 7-methylguanosine caps and 3´ poly-A tails. Prokaryotic
Signals for translation initiation. Initiation sites in prokaryotic mRNAs are characterized by a Shine-Delgarno sequence that precedes the AUG initiation codon. Base pairing between the Shine-Delgarno sequence and a complementary sequence near the 3´
Also Check: What Is The Best Protein To Eat
Iichemical And Physical Properties Of The Amino Acid Glutamine
Glutamine is one of two amino acids in the standard repertoire of 20 for ribosomal protein synthesis that contains a side chain amide group . This imparts a number of interesting properties onto this amino acid. First, the two amide hydrogens on the single, unsubstituted side chain amide nitrogen present the possibility of multiple H bonds. Second, despite the lack of charge in the side chain, a single Gln side chain in isolation is relatively hydrophilic due to its polarity and the ability of the side chain amide to interact with solvent water through hydrogen bonding. In scales of amino acid hydrophobicity, Gin is found with other amino acids whose side chains are compatible with solvent water . In a study of surface accessibility in globular proteins, the vast majority of Gin residues are found to be at least partially surface exposed in the folded tertiary structure . Third, consistent with ChouâFasman and other parameters for predicting secondary structure from amino acid sequence , Gin is about equally likely to be located in α helix, β extended chain, or reverse turns in globular proteins . An interesting and important caveat to the preceding discussion of solubility is that the amino acid Gin is not a particularly soluble amino acid , perhaps due to a strong self-association in the solid state. This self-interaction may in large part account for the tendency of polyGln to aggregate.
Paula G. Vissio, … MarÃa P. Di Yorio, in, 2022
The Three Stages Of Protein Synthesis
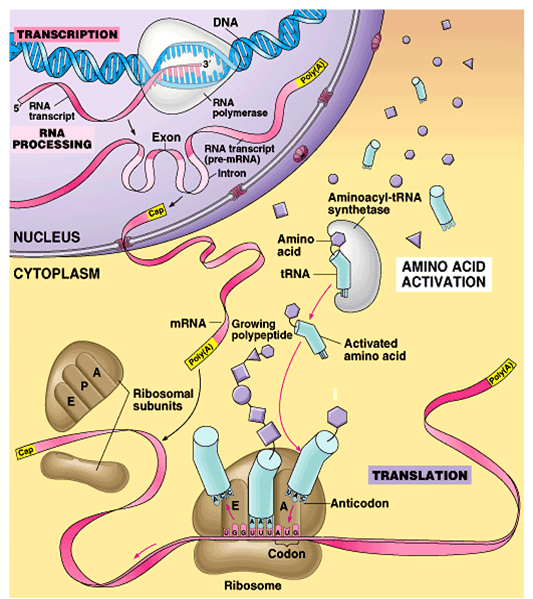
Once mRNA has been transcribed into protein, it can be converted into a functional molecule. The process of decoding these instructions and producing protein is referred to as translation. To make a polypeptide chain, tRNA must be attached to the mRNA molecule and ribosome must be used to construct a polypeptide chain from ribosome. Protein synthesis is halted as a result of this process, which is followed by the release of ribosomes from the host.
Read Also: Best Source Of Protein For Vegetarians
Protein Synthesis Process In Prokaryotes:
Prokaryotes are simple creatures and have only these 5 steps involved in protein synthesis.
- Activation of amino acids
- Transfer of amino acid to tRNA
- Initiation of the polypeptide chain
- Chain Termination and
- Translocation of the protein molecule
Prokaryotes have a single DNA molecule that is used for protein synthesis by transcription and translation. The DNA molecule is so simple that it doesnt even require post-transcriptional and post-translational modifications for its protein synthesis process.
Translocation Of The Protein Molecule:
Two types of polyribosome shave been discovered that are involved in this process:
- Free polyribosomes
- Membrane-bound polyribosomes.
Upon termination of protein synthesis in the free ribosome, the prepared ribosome releases the protein into the cytoplasm. Special types of processes are used to transport some of these specialised proteins to the mitochondria and nucleus.
In membrane-bound polyribosomes, on the other hand, a polypeptide chain that develops on mRNA is introduced into the ER membranes lumen. Some of the proteins also compose parts of the membrane structure.
Even yet, only a few proteins are released into the lumen and integrated into Golgi body vesicles. They can also change the protein through glycosylation, which is the addition of sugar residues. As a result, the vesicles shape a bond with the plasma
membrane and the proteins are sooner or later released.
You May Like: Best Weight Gain Protein Powders
Is Translation Part Of Protein Synthesis
mRNA is read from ribosomes before being translated into amino acid chains that make up the protein, which is synthesized in the cell cytoplasm.
Following DNA transcription, mRNA molecules, which are translated into proteins, are formed during this process. protein is produced by mRNA translation in combination with transfer RNA and ribosomes. During translation, a ribosomal subunit attached to a mRNA molecule. As the ribosome moves through the mRNA molecule, tRNA is released from the P site. When another tRNA recognizes the new mRNA codon, it takes the A binding site back, leaving it open. Molecules of tRNA are released from the complex and attached to each other by new tRNA molecules.
Protein translation is not part of protein synthesis. mRNA is extracted from DNA and then converted into protein. It is present in the nucleus of the cell and is controlled by genes. Following the development of the mRNA, it is read and formed into amino acids at ribosomes. This is referred to as a translation process. Protein synthesis is critical to ribosomes because they serve as the building blocks of protein. They are proteins-rich particles that sit on the surface of ribosomes and interact with mRNA. The ribosomes play an important role in the process of protein chain synthesis. The mRNA is read, and a chain of amino acids is formed.
What Are The Steps Of Protein Synthesis
The three steps are as follows: initiation, extension, and termination. Following the completion of the mRNA processing process, instructions are sent to the cytoplasm of a ribosome.
Protein Synthesis: The Final Step
When mRNA strands are completed, they are sent to the cytoplasm for transcription. The amino acid assembly of the protein is the final step in its synthesis.
Recommended Reading: Best Protein Sources For Vegetarian
How Is Dna Translated Into Protein
Basically, a gene is used to build a protein in a two-step process:
What is DNA translation process?
Translation is the process that takes the information passed from DNA as messenger RNA and turns it into a series of amino acids bound together with peptide bonds. It is essentially a translation from one code to another code .
There Are Many Steps From Dna To Protein
We have seen so far in this chapter that many different types of chemical reactions are required to produce a properly folded from the information contained in a . The final level of a properly folded protein in a cell therefore depends upon the efficiency with which each of the many steps is performed.
You May Like: Whey Protein For Weight Gain
What Are The Initial Steps In Accessing Genetic Information
Nature EducationtranscriptionRNA polymerases
RNA molecules differ from DNA molecules in several important ways: They are single stranded rather than double stranded their sugar component is a ribose rather than a deoxyribose and they include uracil nucleotides rather than thymine nucleotides . Also, because they are single strands, RNA molecules don’t form helices rather, they fold into complex structures that are stabilized by internal complementary base-pairing.
Messenger RNA ribosomal RNA transfer RNA
mRNA is the most variable class of RNA, and there are literally thousands of different mRNA molecules present in a cell at any given time. Some mRNA molecules are abundant, numbering in the hundreds or thousands, as is often true of transcripts encoding structural proteins. Other mRNAs are quite rare, with perhaps only a single copy present, as is sometimes the case for transcripts that encode signaling proteins. mRNAs also vary in how long-lived they are. In eukaryotes, transcripts for structural proteins may remain intact for over ten hours, whereas transcripts for signaling proteins may be degraded in less than ten minutes.
New Roles For Ribosomes

The average mammalian cell contains more than ten million ribosomes. Cancer cells can produce up to 7,500 ribosomal subunits every minute. As a polypeptide-producing factory, the existence, development, and function of every living organism depends on the ribosome.
It was previously thought that eukaryotic ribosomes only played effector roles in protein synthesis . However, recent research now shows that ribosomes also regulate the translation process. They play a part in deciding which proteins are manufactured and in what quantities. The success and results of translation depend on more than the availability of free amino acids and enzymes they also depend on the quality of the ribosomes.
You May Like: How To Get Enough Protein
What Are The Products Of Transcription
The majority of the coding genes are transcribed into messenger RNA. There are some exceptions, and some of the coding genes are transcribed into different types of RNA molecules that carry out quite different functions in the cell. Such types of RNA are ribosomal RNA and transfer RNA they are both required for the successful completion of the protein synthesis process.
The Process Of Translation
Translation is generally divided into three stages: initiation, elongation, and termination . In both prokaryotes and eukaryotes the first step of the initiation stage is the binding of a specific initiator methionyl tRNA and the mRNA to the small ribosomal subunit. The large ribosomal subunit then joins the complex, forming a functional ribosome on which elongation of the chain proceeds. A number of specific nonribosomal are also required for the various stages of the process .
Translation Factors.
The first step in bacteria is the binding of three initiation factors to the 30S ribosomal subunit . The mRNA and initiator N-formylmethionyl tRNA then join the complex, with IF-2 specifically recognizing the initiator tRNA. IF-3 is then released, allowing a 50S ribosomal subunit to associate with the complex. This association triggers the hydrolysis of GTP bound to IF-2, which leads to the release of IF-1 and IF-2 . The result is the formation of a 70S initiation complex that is ready to begin formation during the elongation stage of translation.
Initiation of translation in bacteria. Three initiation factors first bind to the 30S ribosomal subunit. This step is followed by binding of the mRNA and the initiator N-formylmethionyl tRNA, which is recognized by IF-2 bound
Don’t Miss: Whey And Casein Protein Powder
What Are The Steps Of Dna Transcription
Transcription occurs in the three stepsinitiation, elongation, and terminationall shown here.The steps are illustrated in Figure 2.
- Step 1: Initiation. Initiation is the beginning of transcription.
- Step 2: Elongation. Elongation is the addition of nucleotides to the mRNA strand.
- Step 3: Termination.
What is the first step in protein translation?
TranscriptionThe first step of protein synthesis is called Transcription. It occurs in the nucleus. During transcription, mRNA transcribes DNA. DNA is unzipped and the mRNA strand copies a strand of DNA.
Editing By Rna Synthetases Ensures Accuracy
Several mechanisms working together ensure that the synthetase links the correct to each tRNA. The synthetase must first select the correct amino acid, and most do so by a two-step mechanism. First, the correct amino acid has the highest affinity for the active-site pocket of its synthetase and is therefore favored over the other 19. In particular, amino acids larger than the correct one are effectively excluded from the . However, accurate discrimination between two similar amino acids, such as isoleucine and valine , is very difficult to achieve by a one-step recognition mechanism. A second discrimination step occurs after the amino acid has been covalently linked to AMP . When tRNA binds the synthetase, it forces the amino acid into a second pocket in the synthetase, the precise dimensions of which exclude the correct amino acid but allow access by closely related amino acids. Once an amino acid enters this editing pocket, it is hydrolyzed from the AMP and released from the . This hydrolytic editing, which is analogous to the editing by polymerases , raises the overall accuracy of tRNA charging to approximately one mistake in 40,000 couplings.
Hydrolytic editing. tRNA synthetases remove their own coupling errors through hydrolytic editing of incorrectly attached amino acids. As described in the text, the correct amino acid is rejected by the editing site. The error-correction process
Read Also: Isopure Protein Powder Zero Carb
Study Reveals Key Step In Protein Synthesis
Protein-building ribosome caught in the act, showing a crucial mechanism in the translation of genetic code into protein structures
This image of the molecules involved in translation of genes into proteins shows the ribosome in transparent rendering. In the foreground are the messenger RNA , the elongation factor EF-G , and the four sequential positions of transfer RNA as it moves from right to left during translocation . The tRNA positions are like four frames of a molecular movie describing its movement through the ribosome during protein synthesis.
Scientists at the University of California, Santa Cruz, have trapped the ribosome, a protein-building molecular machine essential to all life, in a key transitional state that has long eluded researchers. Now, for the first time, scientists can see how the ribosome performs the precise mechanical movements needed to translate genetic code into proteins without making mistakes.
“This is something that the whole field has been pursuing for the past decade,” said Harry Noller, Sinsheimer Professor of Molecular Biology at UC Santa Cruz. “We’ve trapped the ribosome in the middle of its movement during translocation, which is the most interesting, profound, and complex thing the ribosome does.”
Noller has spent decades working to understand how the ribosome works. Being able to see how it moves, he said, is an exciting moment.
How Is Dna Protein Made
Proteins are the key working molecules and building blocks in all cells. They are produced in a similar two-step process in all organisms called protein synthesis DNA is first transcribed into RNA, then RNA is translated into protein.
What is transcription and translation in DNA?
The process by which DNA is copied to RNA is called transcription, and that by which RNA is used to produce proteins is called translation.
Don’t Miss: High Fat No Protein Foods
Proteins Are Made On Polyribosomes
The synthesis of most molecules takes between 20 seconds and several minutes. But even during this very short period, multiple initiations usually take place on each being translated. As soon as the preceding has translated enough of the sequence to move out of the way, the 5 end of the mRNA is threaded into a new ribosome. The mRNA molecules being translated are therefore usually found in the form of polyribosomes , large cytoplasmic assemblies made up of several ribosomes spaced as close as 80 nucleotides apart along a single mRNA molecule . These multiple initiations mean that many more protein molecules can be made in a given time than would be possible if each had to be completed before the next could start.
A polyribosome. Schematic drawing showing how a series of ribosomes can simultaneously translate the same eucaryotic mRNA molecule. Electron micrograph of a polyribosome from a eucaryotic cell.